研究成果 Research Results
- TOP
- News
- Research Results
- Next-generation test for detecting bacterial endotoxin
Next-generation test for detecting bacterial endotoxin
New advances could help reduce the use of endangered horseshoe crabs for detecting lipopolysaccharide 2020.09.04Research ResultsLife & HealthPhysics & Chemistry
Proteins developed at Kyushu University may soon make tests for identifying the presence of particular endotoxins on bacteria easier to manufacture by ending the dependence on substances from horseshoe crabs.
Found in the outer layer of a number of bacteria, lipopolysaccharide is an endotoxin that can trigger inflammatory responses leading to a condition called sepsis, which can be especially life-threatening in intensive care units. To monitor for contamination of lipopolysaccharide in non-oral drugs, medical devices, and biologics, testing for the molecules has been increasing. However, current tests depend on raw materials from the limited natural resource of horseshoe crabs.
Named after the species name of the American horseshoe crab, the Limulus test uses substances produced by breaking down horseshoe crab amebocytes—the equivalent to mammalian white blood cells—to detect lipopolysaccharide. In response to a minute quantity of bacterial lipopolysaccharide, three precursors of enzymes—referred to as ProC, ProB and ProCE—found in the broken-down cell material cause a chain reaction to form a clot, indicating the presence of the endotoxin.
As an alternative approach, the research team led by Shun-ichiro Kawabata, professor of Kyushu University’s Faculty of Science, has been developing a next-generation Limulus test using recombinant proteins that can be produced from cells grown in the lab as a substitute for the three enzyme precursors. While they have previously made functional recombinant proteins of two of the components, they now report the successful preparation of an alternative for the remaining component, ProCE.
The protein was found to greatly promote the coagulation process and gives the researchers a complete set of alternatives for ProC, ProB and ProCE that could be applied to a next-generation Limulus test to contribute to the development of a very sensitive mixture for the detection of lipopolysaccharide for a variety of biomedical uses without depending on horseshoe crabs.
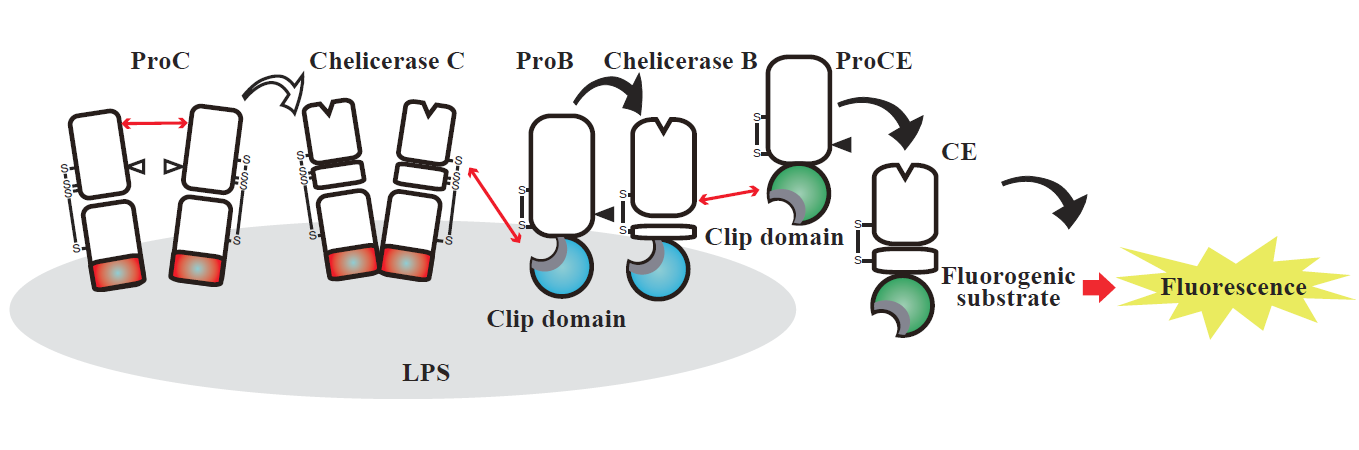
Mechanism for detecting lipopolysaccharide. ProC interacts with lipopolysaccharide through the region colored in red to form a complex between ProC molecules and ProC is converted to the active form Chelicerase C. Then, ProB is recruited to lipopolysaccharide through the region colored in blue (clip domain) and ProB is activated to Chelicerase B by Chelicerase C. The region of ProCE colored in green (clip domain) accelerates the activation of ProCE to the active form CE by Chelicerase B. Finally, the activity of CE is monitored by a synthetic fluorogenic substrate quantitatively to detect lipopolysaccharide.
###
For more information about this research, see “Roles of the clip domains of two protease zymogens in the coagulation cascade in horseshoe crabs,” Keisuke Yamashita, Toshio Shibata, Toshiaki Takahashi, Yuki Kobayashi, and Shun-ichiro Kawabata, The Journal of Biological Chemistry, 295, 8857–8866 (2020). https://doi.org/10.1074/jbc.RA119.012452
Research-related inquiries
Shun-ichiro Kawabata, Professor
Department of Biology, Faculty of Science
Contact information can also be found in the full release.
- TOP
- News
- Research Results
- Next-generation test for detecting bacterial endotoxin