研究成果 Research Results
- TOP
- News
- Research Results
- Sulfur-deficiency-induced repressor proteins optimize glucosinolate biosynthesis in plants
Sulfur-deficiency-induced repressor proteins optimize glucosinolate biosynthesis in plants
2016.10.11Research ResultsLife & Health
Plants use sulfate as a primary sulfur source to synthesize a wide array of metabolites, including the S-containing amino acids Cysteine and Methionine which are incorporated into proteins. Plants of the Brassicaceae family such as oilseed rape, cabbage, broccoli and the model plant Arabidopsis, use sulfate to produce sulfur-rich secondary metabolites, glucosinolates (GSLs) as well as proteins. GSLs are important defense compounds against pathogens and herbivores and provide potential health benefits to human health and nutrition. Under optimum growth condition, when sulfur contents of the environment is sufficient (+S), these metabolites are actively synthesized via the control of several MYB transcription factors. When sulfur content of the environment is limiting (−S), sulfur deficiency Induced genes (SDI1 and SDI2) are strongly induced and acts as negative regulators of GSL biosynthesis. This regulation occurs via protein-protein interaction between SDI1 and MYB28, the major regulator of the pathway. With this mechanism plants can prioritize sulfate usage for primary metabolites and proteins which may well explain the prioritization of growth over defense by plants grown under sulfur deficient environment. This in hand it might be possible to control secondary metabolite contents of medicinal plants through either sulfate fertilization by novel breeding strategies.
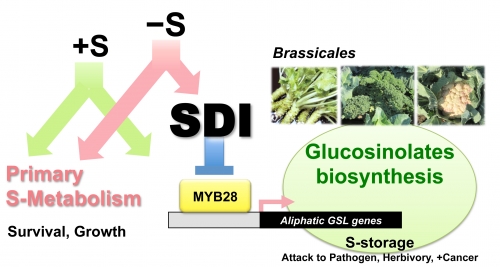
SDI represses biosynthesis of glucosinolates under sulfur deficient environment (−S) by interacting with MYB28, a major regulator of the pathway, which enables plants to prioritize sulfur usage for primary metabolism under −S.
Journal Reference
Sulfur deficiency–induced repressor proteins optimize glucosinolate biosynthesis in plants, ,Science Advances, 10.1126/sciadv.1601087Research-related inquiries
- TOP
- News
- Research Results
- Sulfur-deficiency-induced repressor proteins optimize glucosinolate biosynthesis in plants