研究成果 Research Results
- TOP
- News
- Research Results
- H3mm7, a histone H3 variant, fine-tunes muscle regeneration
H3mm7, a histone H3 variant, fine-tunes muscle regeneration
2018.04.09Research ResultsLife & Health
Genomic DNA in eukaryotes is stored in nuclei as a highly packed structural complex called chromatin. The basic unit of chromatin is the nucleosome in which DNA is wrapped around core histone proteins. Each histone protein is known to have several variants based on amino acid substitutions. In 2015, the research group of Professor Yasuyuki Ohkawa has reported novel mouse H3 variant genes which encoded highly conserved amino acid sequences compared to well-known histones. However, the functions of each histone variant are still unclear because of the diverse nature of histone variants.
In the present study, they have shown for the first time that H3mm7, one of the histone variants, is substantially expressed in muscle stem cells. Furthermore, they found that the loss of the H3mm7 gene resulted in delayed muscle regeneration. Additional experiments revealed that histone H3mm7 has a function to accelerate the gene expression level of muscle-related genes by relaxing the DNA structure in muscle stem cells. This study suggests that histone variants might be another vital factor as gene expression switches in determining unique expression profiles for specific tissues.
This study was supported by JST CREST Grant Number JPMJCR16G1 and JSPS KAKENHI (#25116010, #17H03608, #15K18457, #16K18479,) The research achievement will be published online in Nature Communications on Apr 11, 2018.
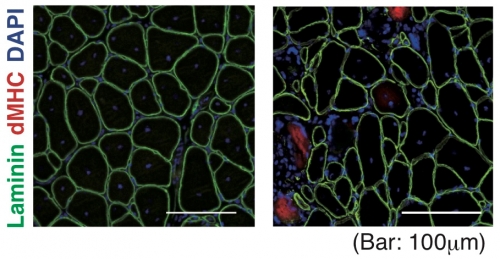
Normal (Left) H3mm7 knockout(Right)
Fig. 1.
Some of the fibers showed incomplete muscle regeneration in H3mm7 knockout mice. Immunostaining of a regeneration marker (dMHC; red) and laminin (green) in Tibialis Anterior muscle at 14 days after muscle injury with cardiotoxin injection is shown.
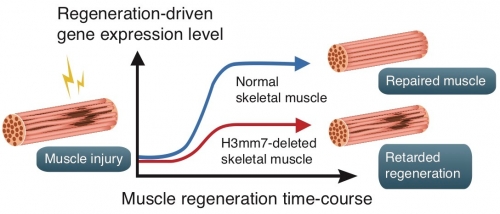
Fig. 2.
Acute injury-induced skeletal muscle regeneration is completed so that muscle stem cells differentiate into mature muscle through the upregulation of the expression of muscle-related genes. However, H3mm7 knockout mice demonstrated impaired regeneration due to the lack of the H3mm7 function to accelerate the expression of muscle-related genes.
Journal Reference
Histone H3.3 sub-variant H3mm7 is required for normal skeletal muscle regeneration, ,Nature Communications , 10.1038/s41467-018-03845-1Research-related inquiries
- TOP
- News
- Research Results
- H3mm7, a histone H3 variant, fine-tunes muscle regeneration